Can You Imagine Why Microbes Produce Chemicals That Are Inhibitory to Other Microbes?
This department describes mutual antibiotic resistance mechanisms in bacteria.
Antibiotics disrupt essential structures or processes in bacteria. This in turn either kills the bacteria or stops them from multiplying. Bacteria have in turn evolved many antibiotic resistance mechanisms to withstand the actions of antibiotics.
How bacteria resist antibiotics
There are two main means for leaner to withstand the effects of an antibiotic:
- To stop the antibody from reaching its target at a high enough concentration
- To modify or bypass the target that the antibiotic acts on
Over time bacteria have evolved many different antibody resistance strategies to attain this.
Antibody resistance mechanisms
one. Stop the antibiotic from reaching its target:
- Pump the antibody out from the bacterial prison cell. Bacteria can produce pumps that sit in their membrane or cell wall. These and so-called efflux pumps are very common in bacteria and can ship a variety of compounds such as point molecules and nutrients. Some of these pumps can also transport antibiotics out from the bacterium, in this style lowering the antibiotic concentration inside the bacterial cell. In some cases mutations in the bacterial DNA can brand the leaner produce more of a certain pump, which in turn increases resistance.
- Subtract permeability of the membrane that surrounds the bacterial cell. Certain changes in the bacterial membrane go far more difficult to pass through. In this way, less of the antibody gets into the bacteria.
- Destroy the antibiotic. There are bacterial enzymes that tin can inactivate antibiotics. One example is β-lactamase that destroys the active component (the β-lactam ring) of penicillins, extremely important antibiotics for treating human infections. In later years, leaner that produce extended-spectrum β-lactamases, so chosen ESBL-producing bacteria, have get a major problem. They can degrade a wide spectrum of β-lactam antibiotics, sometimes also the last resort drugs bachelor for infections with these bacteria.
- Modify the antibiotic. Bacteria can sometimes produce enzymes that are capable of adding unlike chemical groups to antibiotics. This in plow prohibits binding between the antibiotic and its target in the bacterial prison cell.
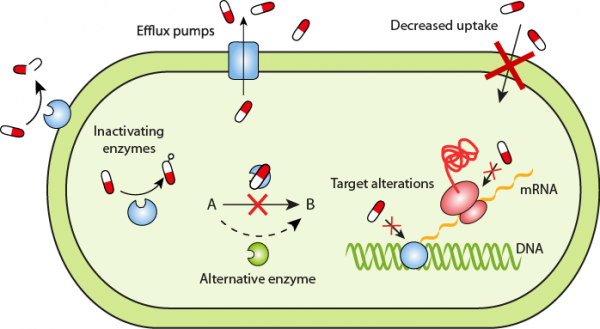
two. Modify or featherbed the target of the antibiotic:
- Camouflage the target. Changes in the composition or construction of the target in the bacterium (resulting from mutations in the bacterial Deoxyribonucleic acid) can stop the antibiotic from interacting with the target. Alternatively, the bacteria tin can add together different chemic groups to the target construction, in this way shielding information technology from the antibiotic.
- Express alternative proteins. Some bacteria are able to produce alternative proteins that can exist used instead of the ones that are inhibited by the antibody. For example, the bacterium Staphylococcus aureus can learn the resistance gene mecA and produce a new penicillin-binding protein. These proteins are needed for bacterial cell wall synthesis and are the targets of β-lactam antibiotics. The new penicillin-bounden protein has low analogousness to β-lactam antibiotics and is thus resistant to the drugs, and the bacteria survive treatment. This type of resistance is the basis in MRSA (methicillin-resistant Staphylococcus aureus).
- Reprogram target. Sometimes bacteria can produce a different variant of a structure it needs. For example, Vancomycin-resistant leaner brand a unlike cell wall compared to susceptible bacteria. The antibiotic is not able to interact every bit well with this blazon of cell wall.
Some bacteria are naturally resistant to certain antibiotics. Imagine for example an antibiotic that destroys the cell wall of the bacteria. If a bacterium does not have a cell wall, the antibiotic will accept no effect. This phenomenon is called intrinsic resistance. When a bacterium that was previously susceptible to an antibiotic evolves resistance information technology is called caused resistance.
Selected Resource
{280251:V8GMEMUX};{280251:EHB3RVFN};{280251:WM2P7FNR};{280251:DQCQ74EJ},{280251:EXZ2BW94};{280251:DQCQ74EJ},{280251:EXZ2BW94} nature default asc no 11299
ane.
Eric Potent. Eric's Medical Lectures. Antibiotic Resistance (Antibiotics - Lecture 9). (2013).
i.
Armando Hasudungan. Microbiology - Leaner Antibiotic Resistance. (2014).
1.
Martínez, J. L. & Baquero, F. Emergence and spread of antibiotic resistance: setting a parameter space. Ups. J. Med. Sci. 119, 68–77 (2014).
1.
Professor Laura Piddock: 'Understanding the basis of antibiotic resistance'. (2014).
1.
Walsh, C. Molecular mechanisms that confer antibacterial drug resistance. Nature 406, 775–781 (2000).
Source: https://www.reactgroup.org/toolbox/understand/antibiotic-resistance/resistance-mechanisms-in-bacteria/
0 Response to "Can You Imagine Why Microbes Produce Chemicals That Are Inhibitory to Other Microbes?"
Post a Comment